Abstract
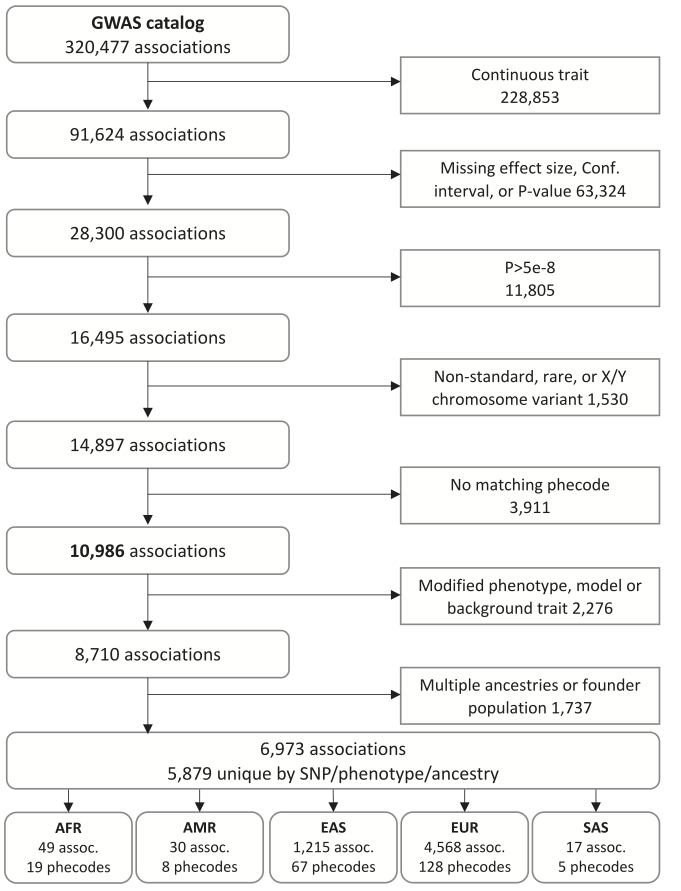
Population-scale biobanks linked to electronic health record data provide vast opportunities to extend our knowledge of human genetics and discover new phenotype-genotype associations. Given their dense phenotype data, biobanks can also facilitate replication studies on a phenome-wide scale. Here, we introduce the phenotype-genotype reference map (PGRM), a set of 5,879 genetic associations from 523 GWAS publications that can be used for high-throughput replication experiments. PGRM phenotypes are standardized as phecodes, ensuring interoperability between biobanks. We applied the PGRM to five ancestry-specific cohorts from four independent biobanks and found evidence of robust replications across a wide array of phenotypes. We show how the PGRM can be used to detect data corruption and to empirically assess parameters for phenome-wide studies. Finally, we use the PGRM to explore factors associated with replicability of GWAS results.
Where applicable, full text and supplement provided for fair use.